This site is being phased out.
Parametrized complexes
Redirect to:
Since Poincare, homology has been used as the main descriptor of the topology of geometric objects. In the classical context, however, all homology classes receive equal attention. Meanwhile, applications of topology in analysis of images and data have to deal with noise and other uncertainty. This uncertainty appears usually in the form of a real valued function defined on the topological space. Persistence is a measure of robustness of the homology classes of the lower level sets of this function.
Since it's unknown beforehand what is or is not noise in the dataset, we need to capture all homology classes including those that may be deemed noise later. In this chapter we introduce an algebraic structure that contains, without duplication, all these classes. Each of them is associated with its persistence and can be removed when the threshold for acceptable noise is set. The last step can be carried out repeatedly in order to find the best possible threshold. The construction follows the approach to analysis of digital images presented earlier.
The topological spaces subject to such analysis are cell complexes. A cell complex is a combinatorial structure that describes how $k$-dimensional cells are attached to each other along $(k-1)$-dimensional cells. Cell complexes come from the following two main sources.
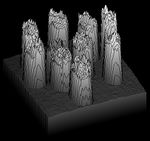
First, a gray scale image is a real-valued function $f$ defined on a rectangle. Given a threshold $r$, the lower level set $f^{-1}((-\infty,r))$ can be thought of as a binary image. Each black pixel of this image is treated as a square cell in the plane. These $2$-dimensional cells have to be combined with their edges ($1$-cells) and vertices ($0$-cells) while in the $n$-dimensional case the image is decomposed into a combination of $0$-, $1$-, ..., $n$-cubes. This process is called thresholding. The result is a cell complex $K$ for each $r$.
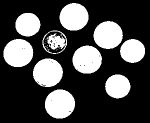
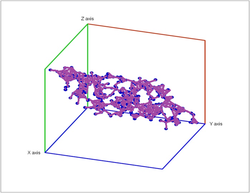
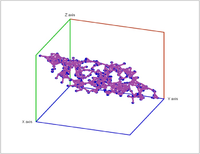
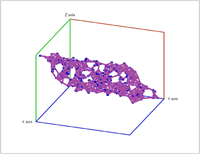
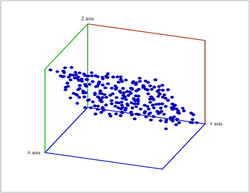
Second, a point cloud is a finite set $S$ in some Euclidean space of dimension $d$. Given a threshold $r$, we deem any two points that lie within $r$ from each other as "close". In this case, this pair of points is connected by an edge. Further, if three points are "close", pairwise, to each other, we add a face spanned by these points. If there are four, we add a tetrahedron, and, finally, any $d+1$ "close" points create a $d$-cell. The process is called the Vietoris-Rips construction. The result is a cell complex $K$ for each $r$.
Next, we would like to quantify the topology of the cell complex $K.$ It is done via the Betti numbers of $K$:
- $B_{0}$ is the number of connected components in $K$;
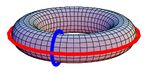
- $B_{1}$ is the number of holes or tunnels ($1$ for letter 'O' or the donut; $2$ for letter 'B' and the torus);
- $B_{2}$ is the number of voids or cavities ($1$ for both the sphere and the torus), etc.
The Betti numbers are computed via homology theory. One starts by considering the collection $C_{k}(K)$ of all formal linear combinations (over a ring $R$) of $k$-cells in $K,$ called $k$-chains.
Combined they form a finitely generated abelian group called the chain complex $C_{k}(K)$, or collectively $C_{\ast}(K).$ A $k$-chain can be recorded as an $N_{k}$-vector, where $N_{k}$ is the total number of $k$-cells in $K$. The boundary of a $k$-chain is the chain comprised of all $(k-1)$-faces of its cells taken with appropriate signs. Then the boundary operator $\partial:C_{k}(K)\rightarrow C_{k-1}(K),k=0,1,...,$ acts on the chain complex and is represented by a $N_{k}\times N_{k-1}$ matrix.
From the chain complex $C_{\ast}(K),$ the homology group is constructed by means of the standard algebraic tools. To capture the topological features one concentrates on cycles, i.e., chains with zero boundary, $\partial A=0$.
Further, one can verify whether two given $k$-cycles $A$ and $B$ are homologous: the difference between them is the boundary of a $(k+1)$-chain $T:A-B=\partial T$ (such as two meridians of the torus). In this case, $A$ and $B$ belong to the same homology class $H=[A]=[B]$.
The totality of these equivalence classes in each dimension $k$ is called the $k$-th homology group $H_{k}(K)$ of $K$, collectively $H_{\ast}(K)$. Then, Betti number $B_{k}$ is the rank of $H_{k}(K)$.
The methods for computing homology groups are well developed. In real-life applications however both digital images and point clouds may be noisy and one needs to evaluate the significance of their homology classes. The approach to this problem has been the following. Instead of using a single threshold and studying a single cell complex, one considers all thresholds and all possible cell complexes.
Since increasing threshold $r$ enlarges the corresponding complex, we have a sequence of complexes: $$ K^{1}\hookrightarrow K^{2}\hookrightarrow K^{3}\hookrightarrow K^{4}% \hookrightarrow\ldots\ \hookrightarrow K^{s},$$ where the arrows represent the inclusions: $$i^{n,n+1}:K^{n}\hookrightarrow K^{n+1}.$$ Let $i^{nm}:K^{n}\hookrightarrow K^{m},n\leq m,$ also be the inclusion. This structure $\{K^{n},i^{nm}\}$ is called a filtration.
We illustrate filtrations on the cell level below.
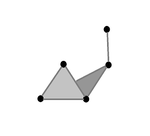
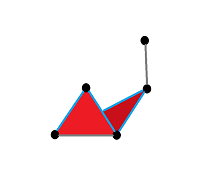
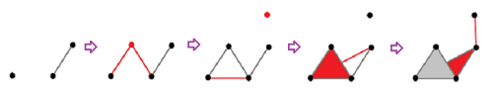
Vietoris-Rips construction:
A digital image constructed pixel by pixel:
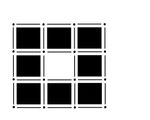
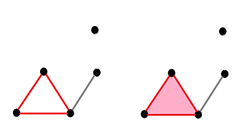
Its homology classes:
Now, each of these inclusions generates a homomorphism $i_{\ast}^{nm}:H_{\ast }(K^{n})\rightarrow H_{\ast}(K^{m})$ called the homology map induced by $i^{nm}.$ As a result, we have a sequence of homology groups connected by these homomorphisms: $$H_{\ast}(K^{1})\rightarrow H_{\ast}(K^{2})\rightarrow\ldots\ \rightarrow H_{\ast}(K^{s})\longrightarrow0. $$ These homomorphisms record how the homology changes as the complex grows at each step. For example, a component appears, grows, and then merges with another one, or a hole is formed, shrinks, and then is filled. We refer to these events as birth and death of the corresponding homology class.
In order to evaluate the robustness of an element of one of these groups the persistence of a homology class is defined as the number of steps in the homology sequence it takes for the class to end at $0.$ In other words,
- persistence = death date - birth date.