This site is being phased out.
Counting fixed and live red blood cells
Fixed red blood cells
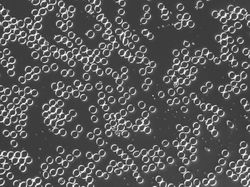
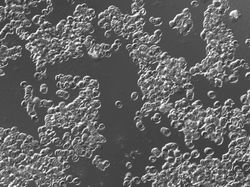
These are fixed red blood cells under microscope. The task is to count them with Pixcavator. They average 7.7 microns in size and were photographed unstained with differential interference contrast lighting. The goal is to count them. I had to crop the image to ensure reasonable processing times.
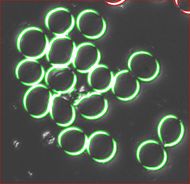
The quality of the image is good, but there is still a problem. Each cell is captured by two light semicircles. These two semicircles aren't connected to each other however (because the light comes from one direction?), so there are no full circles. The result is that cells can't be treated as objects and they aren't captured by the software. Ideally, there should be red contours for each of the cells like in the image on the right.
One way to get around this is to count the semicircles themselves (2 per cell). I ran Pixcavator with the following settings: 1000 for area and 100 for contrast.
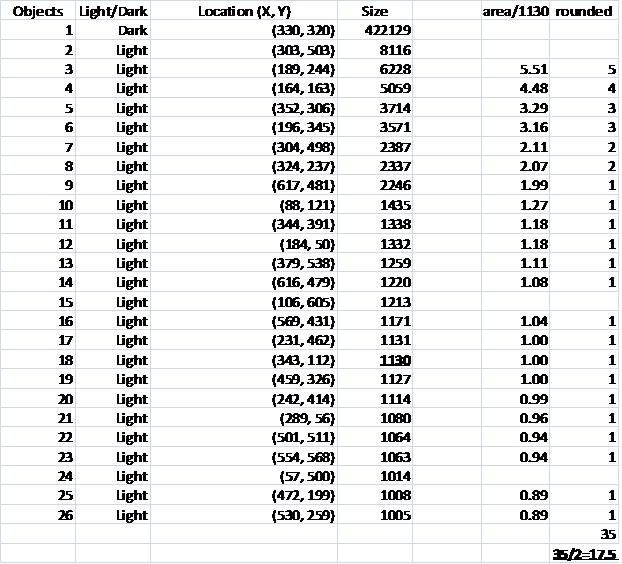
The problem with counting semicircles is that many of some of them touch each other so that they form "clusters". These clusters are what’s captured by Pixcavator. To deal with this problem I used some measurements and some extra computation that followed the analysis. In the last column of the saved spreadsheet (table below) I divided the areas of the clusters of semicircles by the area of one semicircle (“1030”). The total number of semicircles found this way was 35. Then the estimate was 17.5 versus 17 of manual count.
Another way to handle the problem is to start with some preprocessing. Erosion makes the light semicircles grow, they merge and form circular regions. Inside of those lie dark objects captured by Pixcavator. They correspond to cells.
I did 15 rounds of erosion (I had to use Pixcavator’s feature because ImageJ does erosions for binary images only). 15 is a lot as you can see.
Then I analyzed the image with the following settings: contrast 27, saliency 6768. The erosions, however, created several artifacts that had to be unmarked.
This method is more straightforward. With it, however, it is harder to get good results without manual intervention.
Live blood cells
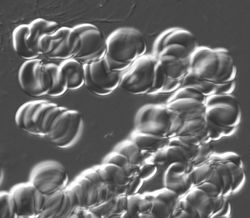
Fixed preparation was fairly easy as the cells were isolated. The living cells tend to adhere and form "rolls".
The image is too messy to analyze, even manually. Best we can do is to evaluate the area covered by these cells.
I had to crop the image to ensure reasonable processing time. To try to estimate the area covered by these cells, I did 7 rounds of erosion. Then I analyzed the image with Pixcavator, settings: area = 67,523. The area of the only dark objects (given by the red contour) was 101,250. Considering 7 rounds of erosion, estimate is a bit off. Then the area covered by the cells is 709x619 (the area of the image) - 101,250 = 337,621 pixels.
Based on this data one may try to estimate the number of cells by dividing the found area by the perceived density of the cells. This number would have to be found manually. Considering the fact that the density varies a lot, the resulting estimate would be quite crude.
This is a clear example of limitations of digital image analysis.
See also Cell counting.
Other image analysis examples